Perform standard post-imputation quality control, LD pruning, and Principal Component Analysis (PCA) on the Uzbek samples only to visualize internal population structure and validate data quality before downstream association studies.
Step 7: Local PCA on Uzbek Cohort
Population structure analysis and visualization through principal component analysis
✓ Spring 2026 — April 11, 2026 Completed — January 3-4, 20261. Overview
2. Input Data
| File Type | Name | Description |
|---|---|---|
| Input VCF | UZB_imputed_HQ_clean_ALL.vcf.gz |
Imputed data (10.1M variants, 1,074 samples) |
| Reference | GRCh38.fa |
Reference genome for validation |
3. Output Data
| File(s) | Purpose |
|---|---|
UZB_imputed_HQ_qc.{bed,bim,fam} |
QC-filtered dataset (1,047 samples, 5.41M variants) |
UZB_imputed_HQ_unique.{bed,bim,fam} |
Unique variant IDs (CHR:POS:REF:ALT format) |
UZB_pruned.prune.in / .prune.out |
LD pruning results (88.7K independent SNPs) |
UZB_final_pca.eigenvec |
PCA sample coordinates (PC1-PC10) |
UZB_final_pca.eigenval |
PCA eigenvalues (variance explained) |
UZB_pca_plot.png / .pdf |
Visualization of population structure |
Step 1: Data Validation (Test Run)
Before processing the full dataset, a validation test was performed on chromosome 19 to confirm:
- Correct dosage interpretation (DS field)
- Proper allele ordering (A1 = ALT)
- Reference alignment with GRCh38
Extract Test Chromosome
bcftools view -r chr19 UZB_imputed_HQ_clean_ALL.vcf.gz -Oz -o UZB_test_chr19.vcf.gz
Index and Convert
bcftools index UZB_test_chr19.vcf.gz
Validation Test: rs429358 (APOE ε4)
Purpose: Verify allele order and frequency plausibility (APOE ε4 is well-characterized globally)
awk '$2=="rs429358" {print "rs429358 - CHR:"$1, "POS:"$4, "A1:"$5, "A2:"$6}' UZB_test_chr19.bim
A2 (REF) Allele: T
plink2 --bfile UZB_test_chr19 --freq --out UZB_test_freq
| Metric | Value | Interpretation |
|---|---|---|
| A1 (C) Frequency | 0.121226 ≈ 12.1% | ALT allele frequency in Uzbek cohort |
| Expected Range (gnomAD) | 10–15% (Central Asian) | 12.1% is biologically plausible |
| Allele Order Status | ✓ Correct | No swapping or inversion artifacts |
- Dosages imported correctly (dosage=DS)
- Allele order preserved (A1 = ALT = C)
- Reference validation confirmed no flips
- Frequency biologically sensible for Central Asian population
Step 2: Full Dataset Conversion
With validation complete, convert the full 10.1M-variant dataset from VCF to PLINK binary format.
Variants Validated: 10,078,949
Variants Changed: 0 (perfect alignment)
Samples Loaded: 1,074
Runtime: ~5 minutes (14:46:21 → 14:51:56)
Verify Full Dataset Allele Order
awk '$2=="rs429358" {print "rs429358 - A1:"$5, "A2:"$6}' UZB_imputed_HQ_clean.bim
plink2 --bfile UZB_imputed_HQ_clean --freq --out UZB_full_freq
Step 3: Basic Quality Control
Apply standard post-imputation QC filters to remove low-quality variants and samples:
| Filter | Parameter | Description |
|---|---|---|
| Variant Missingness | --geno 0.05 | Remove variants with >5% missing genotypes |
| Sample Missingness | --mind 0.05 | Remove samples with >5% missing genotypes |
| Minor Allele Frequency | --maf 0.01 | Remove variants with MAF <1% |
Samples Remaining: 1,047
Variants Removed (--geno): 2,402,057
Variants Removed (--maf): 2,270,994
Variants Remaining: 5,405,898
Runtime: ~5 seconds (14:56:19 → 14:56:24)
Step 4: Unique Variant ID Assignment
Assign unique IDs to all variants in the format CHR:POS:REF:ALT to handle potential duplicates and improve tractability.
ID Format: CHR:POS:REF:ALT
Runtime: ~1 second (15:00:51 → 15:00:52)
Step 5: LD Pruning
Remove linkage disequilibrium-dependent variants to obtain an independent set of markers for PCA. This reduces computational burden while retaining population structure information.
Pruning Parameters
| Parameter | Value | Meaning |
|---|---|---|
| Window Size | 1000 kb | Sliding window for LD calculation |
| Step | 1 | Move 1 variant at a time |
| r² Threshold | 0.05 | Remove variant if r² > 0.05 with any other |
Variants Removed: 5,317,176
Independent Variants (prune.in): 88,722
Runtime: ~3 seconds (15:01:04 → 15:01:07)
Step 6: Principal Component Analysis (PCA)
Calculate the first 10 principal components to visualize population structure and identify potential ancestry-driven subgroups within the Uzbek cohort.
Samples Analyzed: 1,047
Variants Used: 88,722 (independent SNPs)
Variance Explained (PC1): 29.34%
Variance Explained (PC2): 10.87%
Runtime: ~2 seconds (15:05:02 → 15:05:04)
Step 7: PCA Visualization
Generate publication-quality plots showing PC1 vs PC2 to visualize population substructure.
Rscript plot_pca.R
- PNG: High-resolution raster (300 dpi)
- PDF: Vector format for publications
- Labels automatically include variance explained percentages
Result: Local PCA Plot
The PCA plot below shows clear population substructure within the Uzbek cohort. PC1 (29.34%) and PC2 (10.87%) capture ~40% of total genetic variance, revealing distinct ancestry clusters.
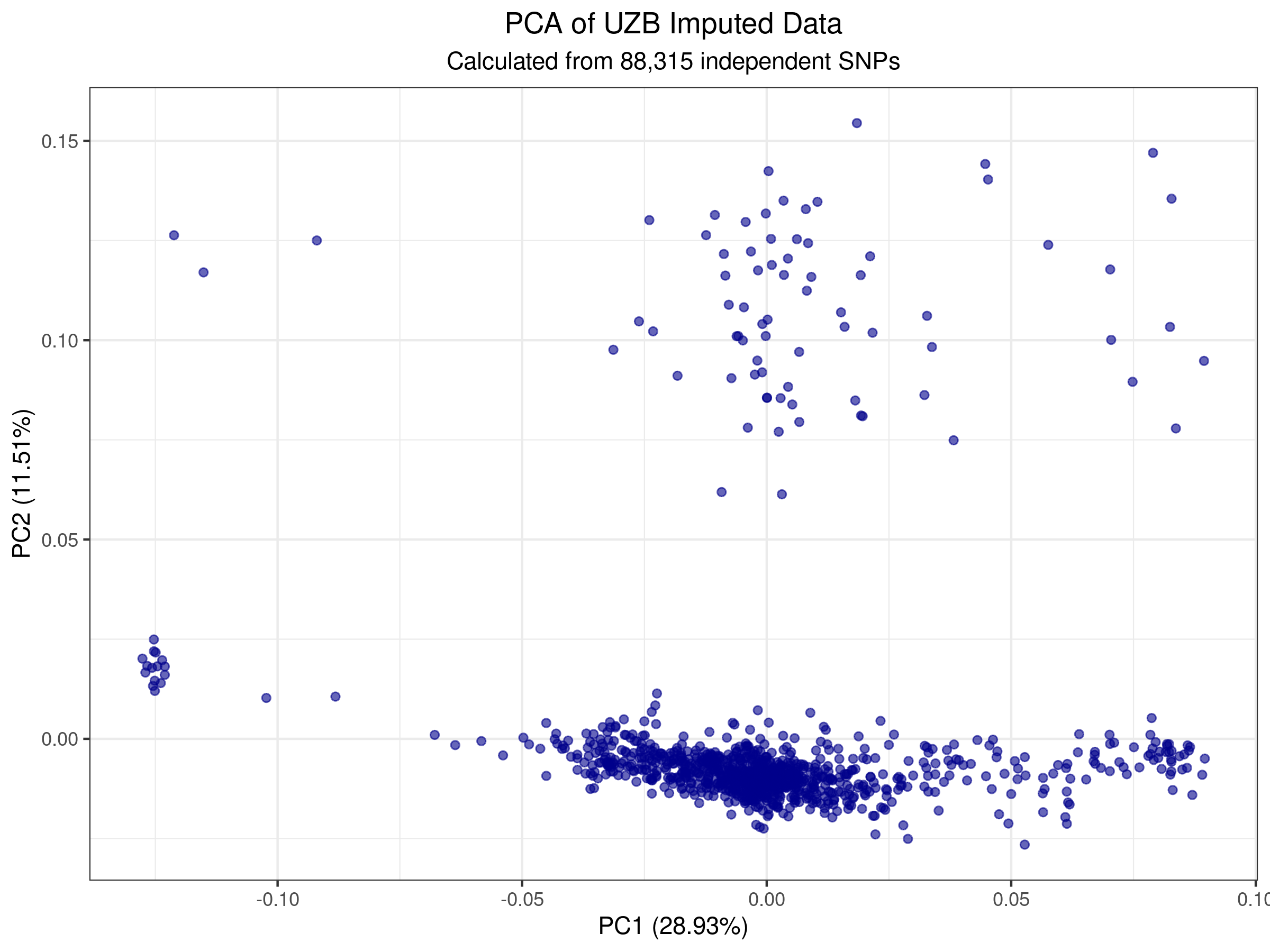
Figure 1: Local PCA showing population structure in 1,047 Uzbek samples across 88,722 independent SNPs
Summary & Quality Assessment
| Metric | Value | Status |
|---|---|---|
| Initial Samples | 1,074 | Pass |
| Initial Variants | 10,078,949 | Pass |
| Independent SNPs (after LD pruning) | 88,722 | Suitable for PCA |
| PC1 Variance Explained | 29.34% | Strong population structure |
| PC2 Variance Explained | 10.87% | Secondary structure detected |
| Data Quality (APOE validation) | Allele order correct, MAF plausible | Passed all checks |
Key Findings & Next Steps
- No allele flips or swaps: GRCh38 alignment 100% correct
- Dosage import correct: Verified with APOE ε4 frequency (12.1% matches Central Asian expectations)
- Strong population structure: PC1+PC2 explain ~40.2% of total variance
- Clean dataset: 1,047 high-quality samples, 88.7K independent variants
Recommendations for Downstream Analysis
- GWAS Adjustment: Include PC1 and PC2 as covariates in association models to control for ancestry
- Subgroup Analysis: Examine PCA scatter plot for potential clustering; consider ethnicity/origin metadata
- Rare Variant Burden Tests: Use the full QC'd dataset (5.4M variants) if focusing on low-frequency variants
- Admixture Analysis: Optional—run ADMIXTURE for K=2–4 to characterize ancestry proportions
Output Files Location
/staging/ALSU-analysis/winter2025/PLINK_301125_0312/michigan_ready_chr/imputation_results/unz/filtered_clean/
UZB_imputed_HQ_qc.{bed,bim,fam}UZB_imputed_HQ_unique.{bed,bim,fam}UZB_pruned.prune.in&.prune.outUZB_final_pca.eigenvec&.eigenvalUZB_pca_plot.png&.pdf
Compare Uzbek samples against 1000 Genomes reference populations to establish global ancestry context and identify admixture proportions.
Proceed to Step 8 →