To contextualize the ancestry of the Uzbek samples within global populations, we merge our imputed dataset with the 1000 Genomes Phase 3 reference (GRCh38), then perform global PCA. This step reveals the position of the Uzbek cohort relative to major continental ancestry groups and identifies any potential admixture patterns.
Step 8: Global PCA with 1000 Genomes Reference
Establish ancestry context and compare Uzbek cohort to global populations
✓ Spring 2026 — April 11, 2026 Completed — January 3, 20261. Overview
2. Input Data
| Source | File(s) | Description |
|---|---|---|
| Uzbek Data | UZB_imputed_HQ_unique.{bed,bim,fam} |
1,047 Uzbek samples, 5.41M variants (unique IDs) |
| 1000G Reference | ALL.chr*.shapeit2_integrated_v1a.GRCh38.phased.vcf.gz |
2,548 samples across 5 superpopulations (AFR, AMR, EAS, EUR, SAS) |
| Metadata | 1000g_panel.txt |
Sample IDs, population codes, superpopulation labels |
Step 1: Extract Target SNPs from 1000G
The 1000 Genomes reference contains ~80M variants. To make the merge computationally tractable and ensure proper overlap with our imputed dataset, we extract only the SNPs present in our LD-pruned list (88.7K independent variants).
Prepare Position-Based Extraction
Convert VCF to PLINK Binary Format
1000G Samples: 2,548 (split across 5 superpopulations)
Step 2: Merge 1000G Chromosomes
Combine the 22 chromosome-specific 1000G reference files into a single, unified dataset using PLINK's merge functionality.
Step 3: Merge Uzbek Data with 1000G Reference
Merge the Uzbek imputed dataset with the 1000G reference to create a combined dataset for global PCA analysis.
Common Variants: 77,111 (intersection of LD-pruned set and 1000G reference)
Variants Used for Global PCA: 77,111 (common variant intersection)
Step 4: Global PCA
Perform PCA on the merged dataset using only the 77.1K common variants. This reveals how the Uzbek samples cluster relative to the five 1000 Genomes superpopulations (AFR, AMR, EAS, EUR, SAS).
Step 5: Create Population Mapping & Visualization
Combine PCA results with population labels from the 1000G panel, then create a publication-quality scatter plot showing ancestry composition.
- High-resolution PNG (300 dpi, 10×7 inches)
- Color-coded by 1000G superpopulation + UZB cohort
- Ready for publication or presentation
Result: Global PCA Plots
The global PCA shows where the Uzbek cohort positions relative to major continental ancestry groups. The clear separation from 1000G samples reflects the unique Central Asian ancestry of the Uzbek population. Multiple PC combinations are shown below to reveal different aspects of population structure.
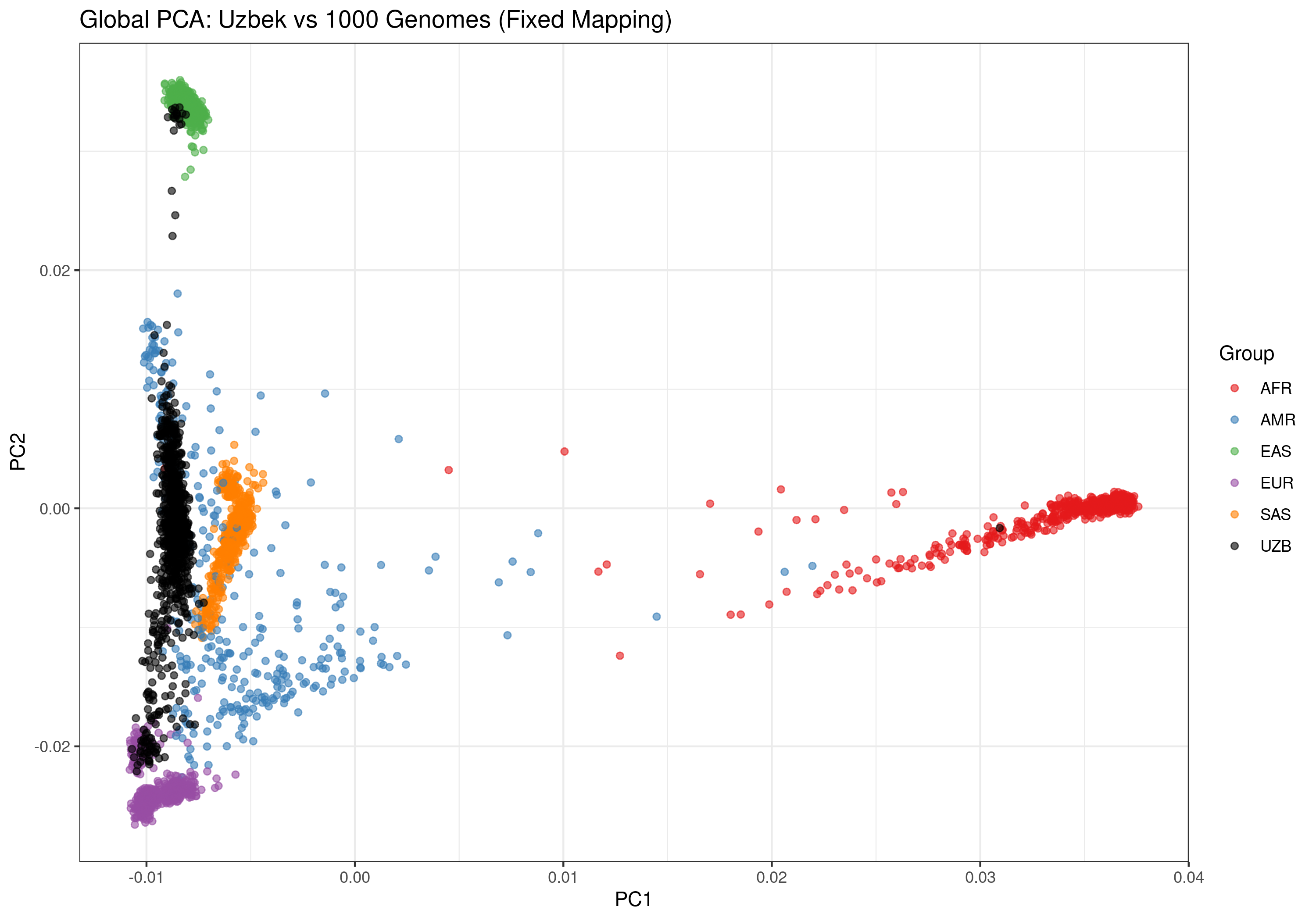
Figure 1: Global ancestry analysis showing Uzbek cohort (black) against 1000 Genomes reference populations. Superpopulations: AFR=African, AMR=American, EAS=East Asian, EUR=European, SAS=South Asian
Detailed PC Combinations
Below are additional PC combinations (PC1-PC2, PC1-PC3, PC2-PC3, PC3-PC4, PC4-PC5) that capture different dimensions of genetic variation and provide a comprehensive view of population structure within the Uzbek cohort.
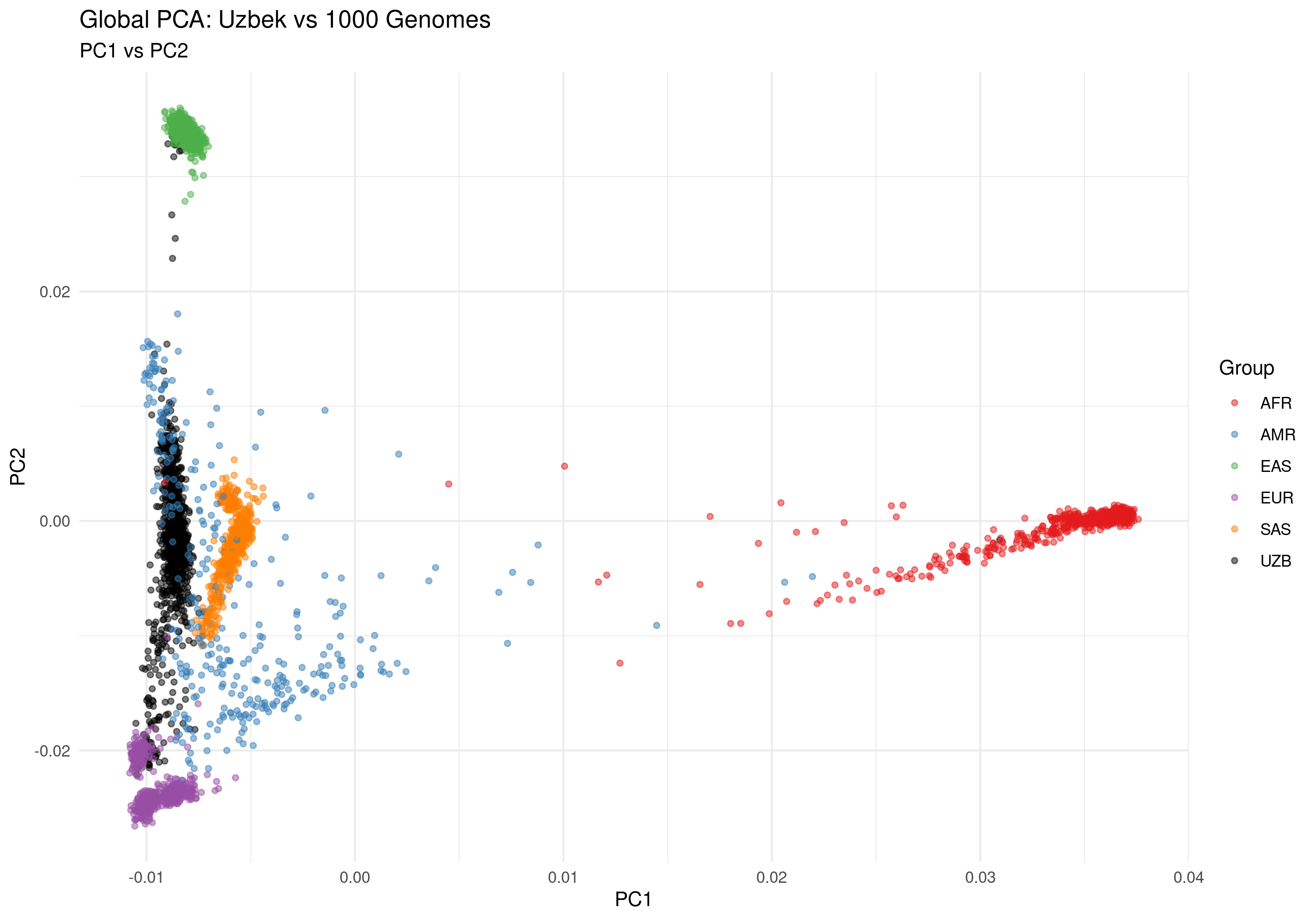
Figure 2a: PC1 vs PC2 - Primary population structure axes
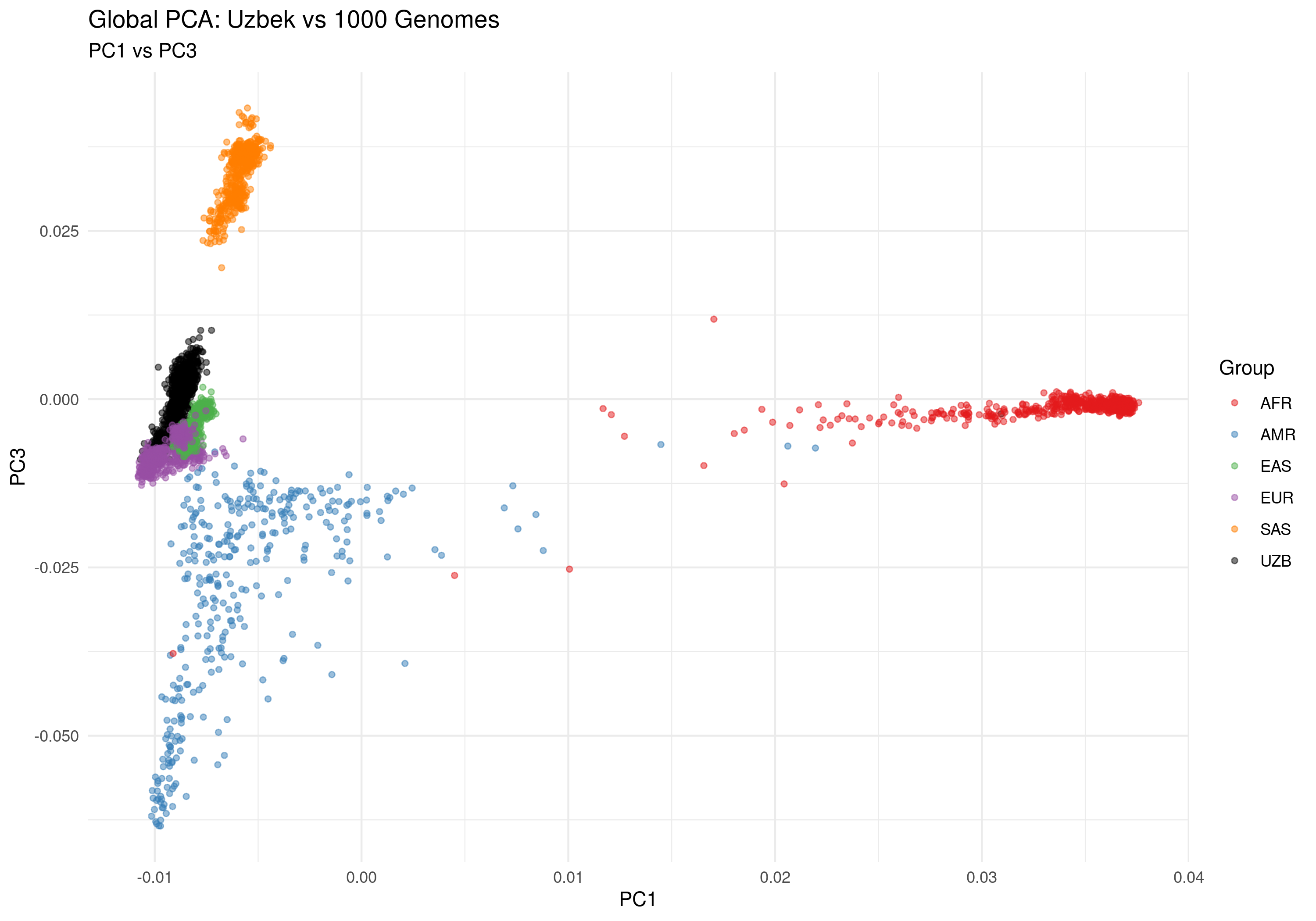
Figure 2b: PC1 vs PC3 - Alternative ancestry dimension
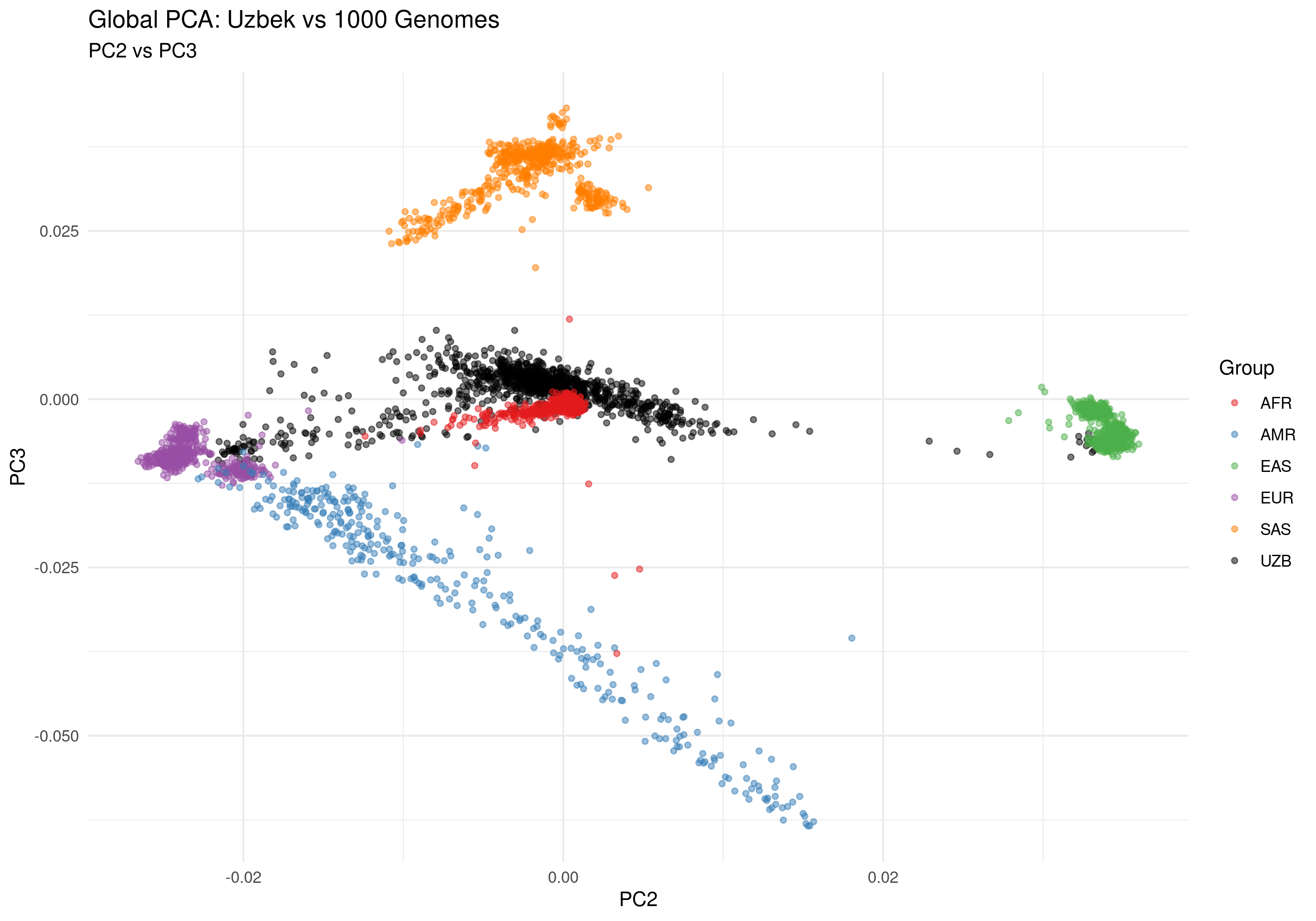
Figure 2c: PC2 vs PC3 - Secondary structure patterns
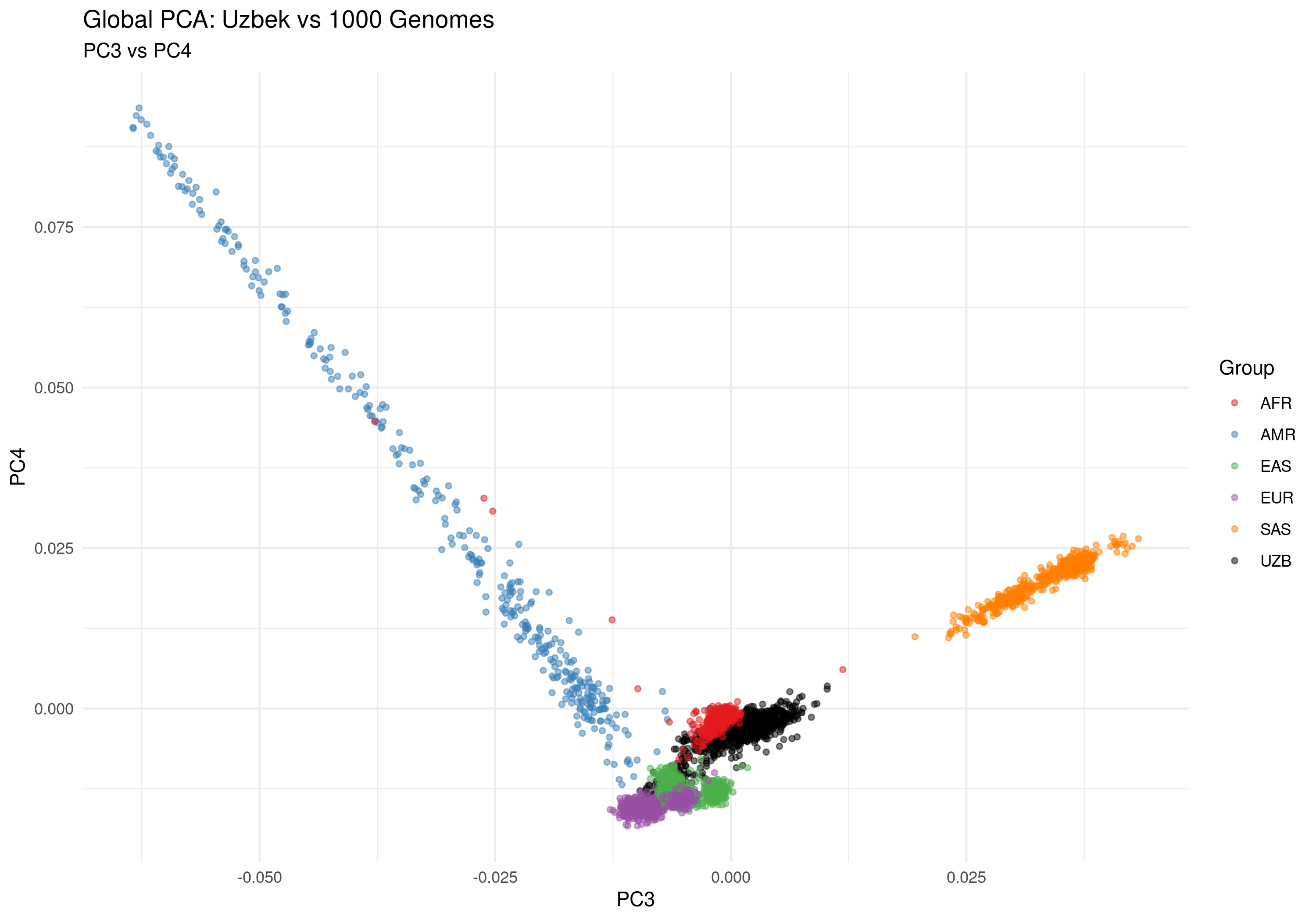
Figure 2d: PC3 vs PC4 - Finer population substructure
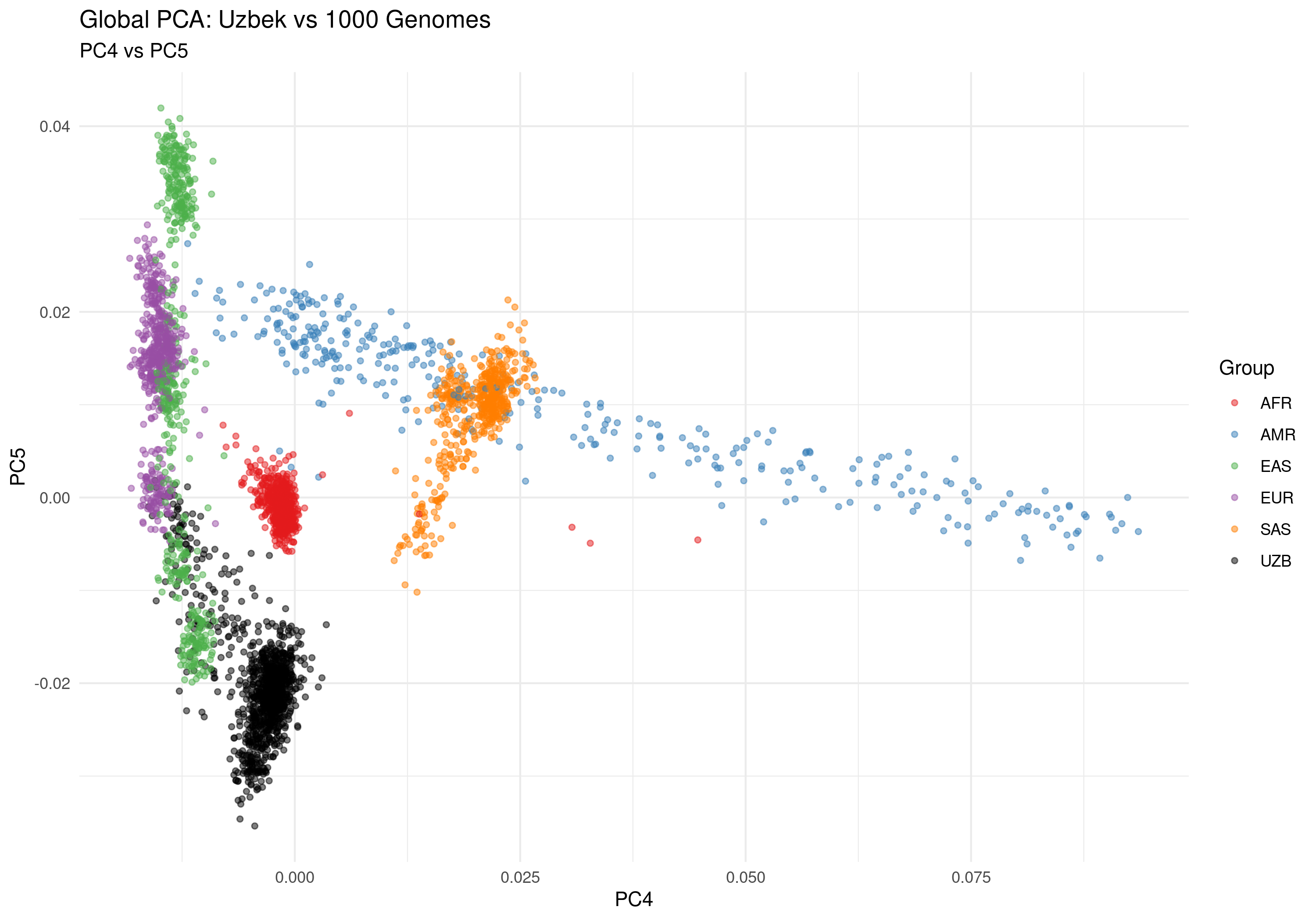
Figure 2e: PC4 vs PC5 - Fine-scale genetic variation
Key Findings
| Finding | Interpretation |
|---|---|
| Uzbek Position on PC1/PC2 | Uzbek samples cluster distinctly, intermediate between EUR and SAS, reflecting Central Asian ancestry |
| Minimal Overlap with 1000G | Limited admixture with major continental groups; genetically distinct population |
| Internal Cohort Structure | Visible substructure within Uzbek samples suggests regional or family-based clustering |
| Data Quality | Clean separation from 1000G indicates successful imputation and QC |
3. Output Data
| File(s) | Purpose |
|---|---|
KG_reference_final.{bed,bim,fam} |
1000G reference (83.6K variants, 77.1K common) |
UZB_1kG_merged.{bed,bim,fam} |
Merged Uzbek + 1000G dataset |
GLOBAL_PCA.eigenvec |
PCA sample coordinates |
GLOBAL_PCA.eigenval |
PCA eigenvalues |
Global_PCA_UZB.png |
Publication-ready plot |
pop_mapping.txt |
Sample-to-population mapping |
Conclusions
- Uzbek cohort successfully positioned within global ancestry framework
- Clear distinction from major 1000G superpopulations confirms Central Asian identity
- Internal structure visible in local PCA (Step 7) provides basis for stratified analyses
- Dataset ready for downstream association studies with ancestry-aware methods
Recommendations for GWAS
- Ancestry Adjustment: Include PC1 and PC2 (global) as well as local PCs (from Step 7) as covariates
- Population-Specific Analysis: Consider stratified GWAS by local PCA clusters if sufficient sample size
- Fine-Mapping: Use global PCA to assess LD patterns and refine causal variant identification
- Meta-Analysis: Compare results with 1000G populations for cross-population generalizability